
Methanogenic Archaea are the only organisms on Earth that can reduce carbon dioxide (CO2) all the way to methane (CH4), and we are fascinated by this unique metabolic pathway. Our model organism for studying the methanogenic Archaea is Methanococcus maripaludis, so named because of its isolation from salt-marsh sediments (-mare, sea; -palus, marsh). M. maripaludis is a great model organism due to its rapid growth in the lab, a complete genome sequence, and a robust set of genetic tools.
Metabolic versatility in hydrogenotrophic methanogens.
Hydrogenotrophic methanogens are those that utilize hydrogen as the primary electron donor for CO2 reduction. However, these organisms possess greater metabolic versatility in that some can additionally use formate, ethanol, secondary alcohols, and possibly carbon monoxide as electron donors for metabolism. We are working to understand how electrons derived from these alternative electron donors feed into each of the four reductive steps of methanogenesis and how genes are regulated in response to the availability of different electron donors.
Biochemistry of heterodisulfide reduction
Soluble heterodisulfide reductase (Hdr) is a flavin-based electron-bifurcating enzyme that couples the first and last steps of methanogenesis. Hdr is a central metabolic hub in methanogenesis and can utilize multiple electron donors for catalysis. We are interested in how electrons enter Hdr through associated hydrogenases or formate dehydrogenases, make their way to the bound flavin in the alpha subunit, and exit the electron bifurcating center.
In addition to the soluble Hdr, members of the Methanosarcinales possess a cytochrome containing, membrane-bound Hdr that accepts electrons from a lipid-soluble phenazine: methanophenazine. While phenazine biosynthesis is well studied in the bacterial domain, little is known of the enzymes responsible for methanophenazine synthesis. We are interested in exploring this pathway using genetic techniques.
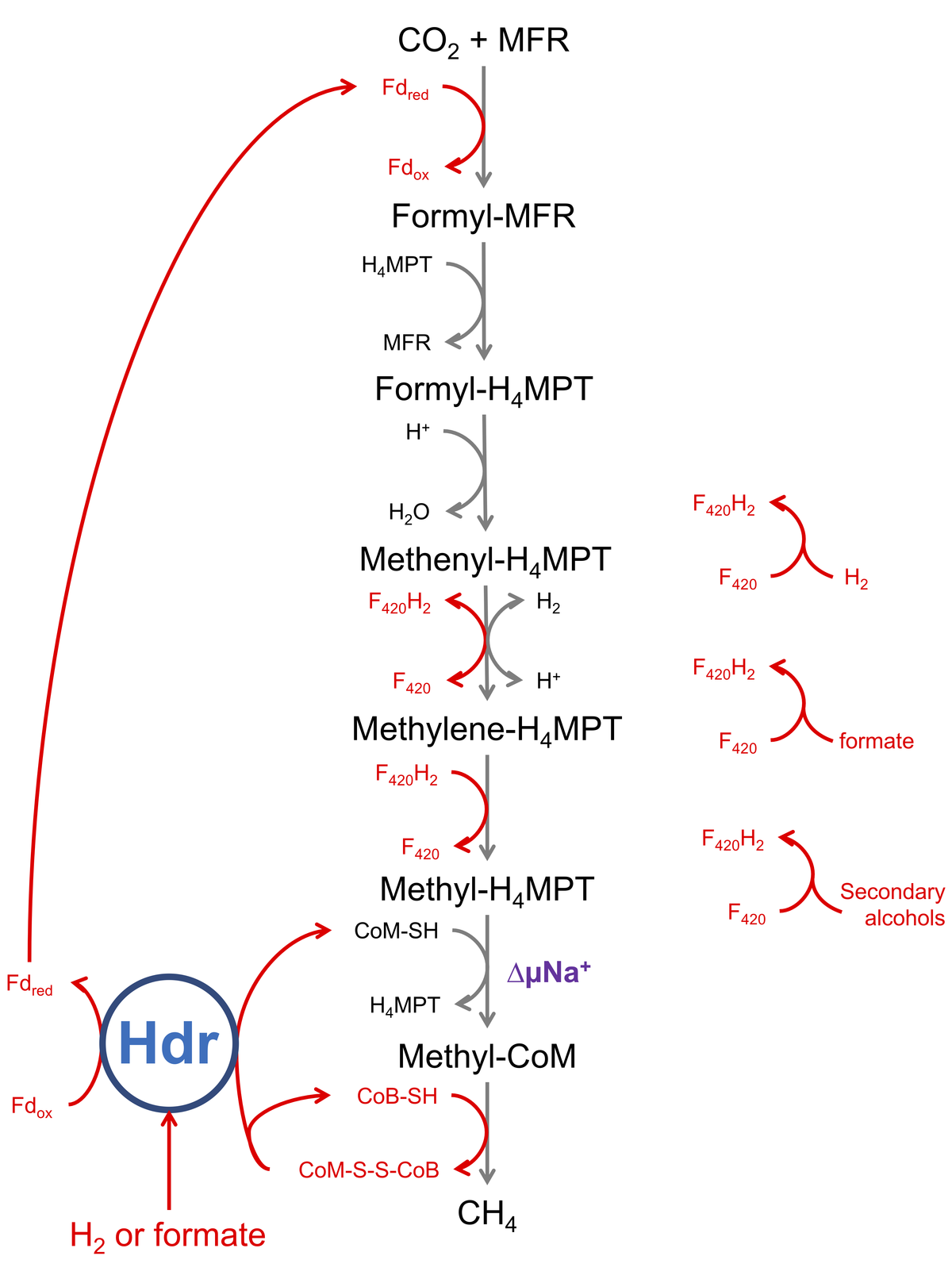
Hydrogenotrophic methanogenesis. Electron transfers are highlighted in red. Hdr is the site of electron bifurcation.
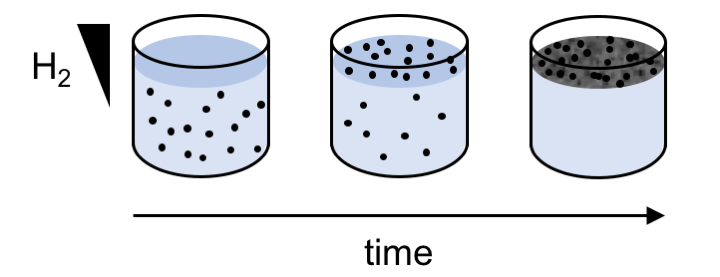
A model of biofilm formation by M. maripaludis at the liquid-atmosphere interface of a standing culture.
Biofilm formation in M. maripaludis
In nature, microbes are primarily found in surface associated communities known as biofilms. However, most microbiological research is performed using planktonic liquid cultures, and the study of anaerobes is no exception. Recent evidence suggests that methanogens prefer to form biofilms near a source of hydrogen. We have developed a high-throughput method to grow M. maripaludis as a biofilm and are beginning to explore the genetic underpinnings of the biofilm lifestyle in this organism.